KU Research Uses Machine Learning to Identify Potential Biomarkers for Respiratory Disease
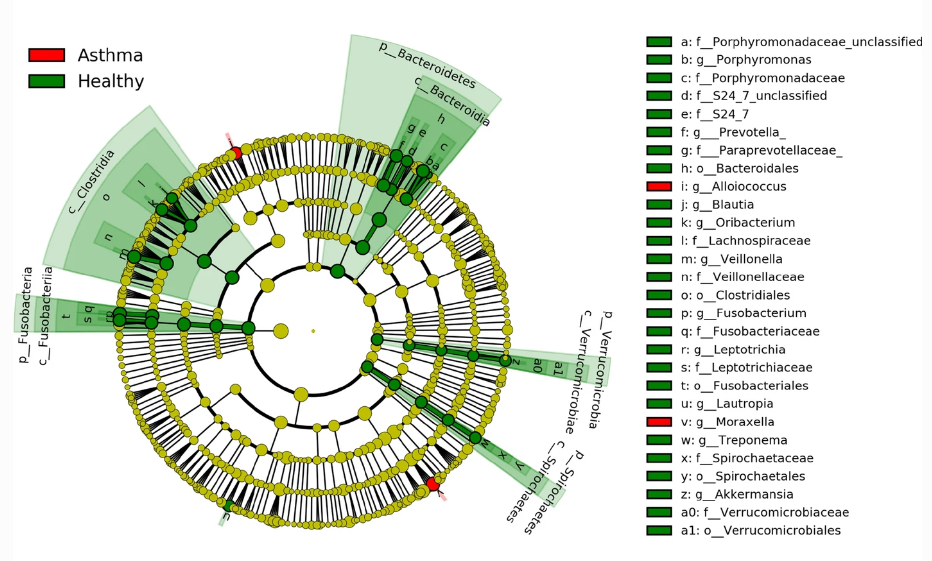
Harnessing the power of machine learning allowed Khalifa University researcher to identify 57 biomarkers for asthma, promising novel avenues for personalized asthma management.
Asthma is a respiratory puzzle that affects over 358 million people globally. Its triggers are vast, ranging from dust and tree pollen to weather and exercise, and its symptoms can escalate from mere coughs to fatal attacks. Asthma is one of the most common non-communicable diseases in adults and the most common chronic ailment in children, but while its genetic links are increasingly being uncovered, the full picture remained elusive.
Now, researchers are turning to the human microbiome, a community of trillions of microbes, encompassing bacteria, fungi, and even viruses, which play pivotal roles in human health and illnesses. While most microbiota studies focus on the gut, newer research is unveiling the profound impact of respiratory microbiota. Illnesses such as asthma, bronchitis, and chronic obstructive pulmonary disease have shown correlations to shifts in these microbial communities.
Dr. Mohammad Bataineh, Assistant Professor of Molecular Biology and Genetics at Khalifa University, used advanced machine learning techniques to characterize the respiratory microbiota in asthmatic patients compared with healthy people and explore the potential of microbiota as a biomarker for asthma diagnosis and prediction. With Reem Salman and Ayman Alzaatreh from the American University of Sharjah, Dr. Bataineh’s research found 57 microbial markers that could be used to characterize notable microbiota composition differences between those with asthma and those without. Their results were published in the Journal of Big Data, a top 1% journal for research in computer science. The research team hope that in an era of precision medicine, their discovery of the respiratory microbiota associated with asthma could lead to valuable applications for individualized asthma care.
“Differences in the microbial populations in the respiratory airways have always been related to the diagnosis of asthma across many studies,” Dr. Bataineh said. “Previous works have shown antibiotic use in early life is associated with the development of childhood asthma, while other studies have also shown that people who grow up in surroundings with high microbiological diversity have a much lower chance of developing asthma. We know that diversified microbial environments can promote diversified human microbiota, leading to a reduced risk of atopic illness and improved lung function, especially in children.”
In the past decade, machine learning has woven its way into biological research, turning vast, intricate datasets into comprehensible patterns. Where traditional methods of analyzing microbial populations might stumble, machine learning algorithms race ahead, adept at navigating the dense forests of high-dimensional data.
The research team applied advanced machine learning techniques to a dataset of 5,853 operational taxonomic units (OTUs) — microbial categories — for 40 patients to identify a collection of the most relevant OTUs. Their results not only corroborated the research that microbiota differences in the respiratory airways are related to asthma diagnosis, but identified 57 microbial biomarkers for diagnosis and therapeutic strategies.
They also noticed variations in microbial abundances across age groups. Pediatric asthmatics, for example, showed significant abundance of Proteobacteria, while other phyla were prominent among adult asthmatics, providing a potential fingerprint for diagnosis.
“Our results demonstrate that the respiratory microbiota’s taxonomic composition and diversity were all significantly different between healthy and asthmatic samples,” Dr. Bataineh said. “We also noted greater diversity and richness of respiratory microbiota in healthy samples compared with asthmatic patients.”
While the study did uncover key respiratory bacterial species associated with asthmatic patients and provide new insights into their role in asthma pathogenesis, there is room for future research. The team recognized that their study was based on a relatively small sample size and a specific type of sample, which makes it challenging to differentiate between the microbiota of the upper and lower respiratory tracts. They also noted the presence of specific bacterial colonies, like Moraxella, can alter the likelihood of viral infections, and emphasized the need for further studies before these biomarkers can be concrete.
Dr. Bataineh’s work forms a foundation for future studies to delve deeper into the relationship between respiratory microbiota and its role in diseases like asthma. The integration of machine learning in the research process has helped pinpoint potential biomarkers, which could pave the way for novel therapeutic strategies targeting the microbiome.
Jade Sterling
Science Writer
6 September 2023